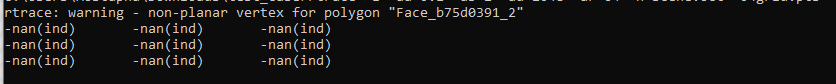
Hi. I have multiple illuminance results showing the same error. When the file is opened in pollination, the entire grid results read as “-nan”. Is there any way to read this file?
Hi @monkspaces, I have to double-check your run to see how it has happened but the error is not with the handler an is actually happening in the simulation. I opened the results file and the results are set to -nan! You can see them here. These values should be numbers.
Hi @mostapha
Could the simulation error be an issue with the modelling?
Thanks for checking this out for me 
Hi @monkspaces, I checked the model and the run and I can confirm that the rtrace command returns -nan (not a number) values.
The model looks fine. Not sure what’s going on when we create the octree and try to run rtrace. I suspected it has to do with the sky but using a different sky didn’t solve the problem. I can render the model using Rhino/Grasshopper which suggests the model itself doesn’t have an issue.
@sarith or @mikkel, can you have a look to this case. Try to run the run.bat file which creates the octree and then runs the rtrace command.
test_case.zip (467.8 KB)
Here is what I get:
So, running rtrace with just the sky works:
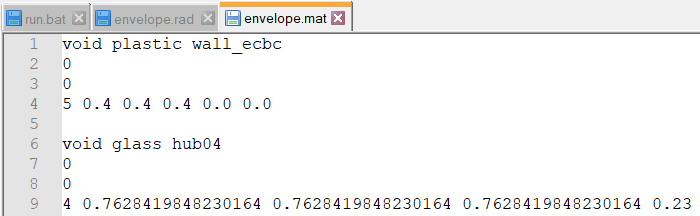
Which means it must be envelope.mat/envelope.rad causing the issue. I looked into envelope.mat:
The refractive index of the glass modifier is set to 0.23. If I change the value to 1.00 or higher it runs:
EDIT: For good measure, @monkspaces, I will just add that the default refractive index of glass in Radiance is 1.52, and a value of 1.00 is that of air.
Hi @mikkel
I ran the model with the default refractive index of the glass. Got results within 5 minutes.
Thank you for helping me figure this out.